2023.03.16 Thursday
Solving the Mystery of Left-Handed Amino Acids in Primordial RNA Reactions
Scientists use computer simulations to investigate chiral selective behavior towards L-alanine in primordial RNA minihelix aminoacylation reactions
The proteins that make up life on earth are synthesized on ribosomes and exclusively composed of L- or left-handed amino acids. A primordial RNA minihelix shows preference for L-amino acid over D-amino acid during its aminoacylation, which is a key reaction determining the chiral selection of amino acids in the evolution of life. However, the mechanism driving such chiral selectivity in RNA is still a mystery. In a new study, two researchers from Tokyo University of Science used quantum mechanics/molecular mechanics-based simulations to decode the physiochemical origin of the chiral selective aminoacylation in the primordial RNA.
While humans have an esthetic liking for symmetry in everything, nature prefers asymmetric, single-handed forms when it comes to amino acids―the building blocks of proteins, and, by extension, all things biological.
But what does the term left-handed amino acids mean? Amino acids are organic molecules with carbon atoms forming their central skeleton. They usually contain a chiral carbon, i.e., a carbon attached to four different functional groups. This makes amino acids optically active chiral molecules with structures that are mirror images of each other and are not superimposable, just like our left and right hands (Fig. 1). Based on the way these molecules interact and steer the direction of light, they are divided into L-type (left-handed) and D-type (right-handed). What's fascinating is that all the proteins produced by nature are made up of L-type amino acid chains. This phenomenon is known as biological or amino acid homochirality, and its evolutionary origin and underlying mechanism have been a long-standing scientific puzzle.
To solve this puzzle, it is important to understand the chiral selectivity seen in certain reactions in primordial RNA minihelices (Fig. 2, upper). So, Associate Professor Tadashi Ando and Professor Koji Tamura from Tokyo University of Science (TUS) investigated the possible mechanisms behind these reactions and their chiral selectivity. Their study published in Life used computer simulations (Fig. 2, lower) to clarify why the amino acid L-alanine was preferred over D-alanine during primordial RNA aminoacylation reactions, without any ribozymes or enzymes steering the selectivity. Aminoacylation reactions are biologically important reactions that involve the attachment of an amino acid to a tRNA during translation. "Many previous studies have suggested that chiral selectivity in aminoacylation might be caused by the steric clash of the amino acid side chain in the constraint of a double helical conformation. However, this has not been fully explained." stated Dr. Ando while highlighting the fact that "This study will help us get closer to understanding how organisms on earth came to utilize L-amino acids and what role they play in the evolutionary continuum."
Direct observation of chiral selective reactions in RNA via experimentation is quite challenging. So, the team adopted a molecular dynamics (MD) simulation strategy that combined the power of quantum mechanical (QM) calculations based on Schrödinger's equation with molecular mechanics (MM) calculations based on Newton's classical mechanics. The team went for the QM/MM-based MD method instead of conventional MD methods because the former can provide atomic-level structural pictures at various stages during the chemical reaction, including those involving bond breaking and formation, unlike the latter.
Once equipped with a robust QM/MM strategy, the team ran simulations of L- and D-alanine aminoacylation reactions on modeled RNA. A free-energy profile analysis revealed that the energy barrier for the L-alanine reaction was 9 kcal/mol lower than that of the D-alanine reaction (Fig. 2, lower). The model mechanisms also showed while in its transition state―i.e, a short-lived molecular configuration where the reaction energy is maximum―L-amino acid had much more electrostatic stability than its D-type counterpart, due to the geometrical arrangement of functional groups in its structure. This observation provided a plausible reason for the previously unexplained trend of selective aminoacylation of L-amino acids in RNA minihelices.
The insights presented in the paper have brought us a step closer to understanding the mechanisms driving the homochirality of amino acids, which is an essential step toward decoding the chemical origin of life. The team believes that in addition to unraveling this mystery, the precise use of QM/MM calculations highlighted in the study will encourage more researchers to use these techniques for further fundamental and application-based research. "tRNA aminoacylation is a key reaction in protein synthesis, and the elucidation of L-amino acid selectivity using computational science is expected to lead to new developments in protein engineering and nucleic acid engineering that control chiral selective reactions" concludes Dr. Ando.
Researchers at TUS are unlocking the universal preference one reaction at a time!
Reference
| Title of original paper | : | Mechanism of Chiral-selective Aminoacylation of an RNA Minihelix Explored by QM/MM Free-Energy Simulations |
| Journal | : | Life |
| DOI | : | 10.3390/life13030722 |
Figure 1
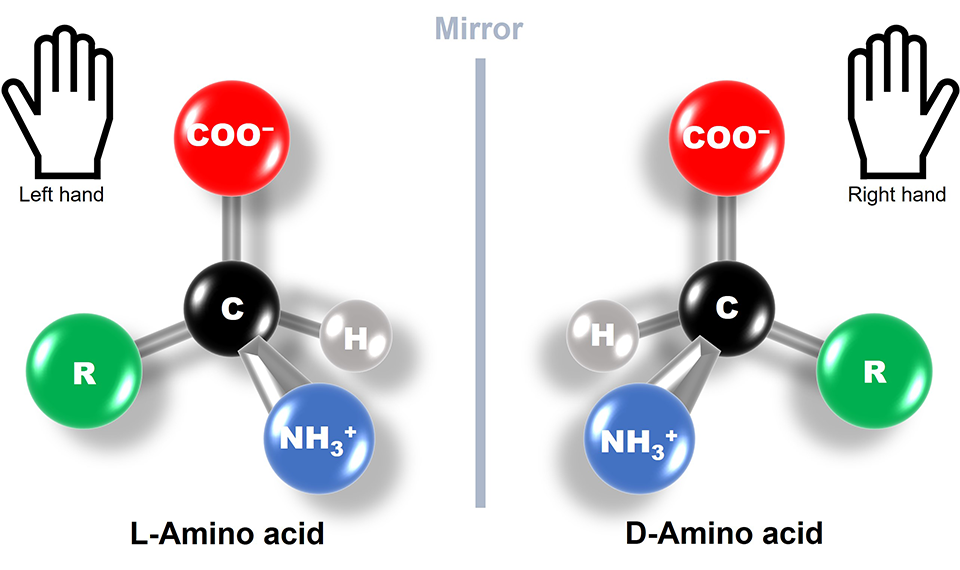
Image caption: Naturally occurring proteins are exclusively left-handed; TUS researchers find out why.
Image title: Amino acids are chiral molecules, i.e., they can be left handed or right handed. However, only L-Amino acids occur in nature.
Image credits: Tadashi Ando from TUS
Image license: Original content
Figure 2
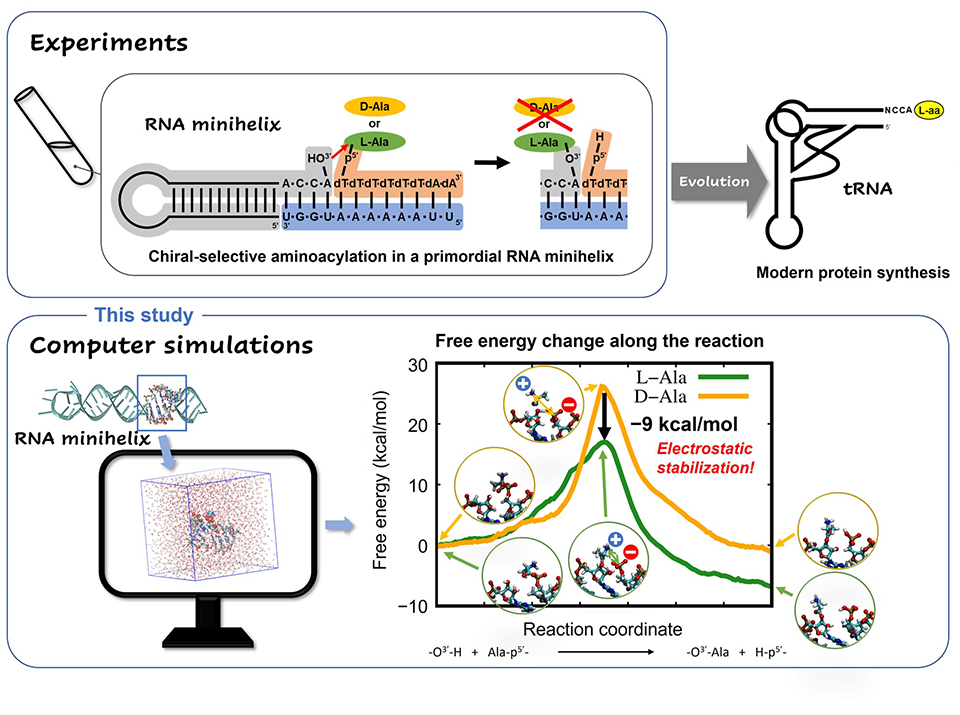
Image caption: Chiral-selective chemical reaction in a primordial RNA minihelix studied using experiments and computer simulations.
Image title: TUS researchers simulate aminoacylation, a reaction that occurs in RNA, to understand why only certain amino acids occur exclusively in nature.
Image credits: Tadashi Ando from TUS
Image license: Original content
About The Tokyo University of Science
Tokyo University of Science (TUS) is a well-known and respected university, and the largest science-specialized private research university in Japan, with four campuses in central Tokyo and its suburbs and in Hokkaido. Established in 1881, the university has continually contributed to Japan's development in science through inculcating the love for science in researchers, technicians, and educators.
With a mission of "Creating science and technology for the harmonious development of nature, human beings, and society," TUS has undertaken a wide range of research from basic to applied science. TUS has embraced a multidisciplinary approach to research and undertaken intensive study in some of today's most vital fields. TUS is a meritocracy where the best in science is recognized and nurtured. It is the only private university in Japan that has produced a Nobel Prize winner and the only private university in Asia to produce Nobel Prize winners within the natural sciences field.
■
Tokyo University of Science(About TUS)

About Associate Professor Tadashi Ando
from Tokyo University of Science
Tadashi Ando is currently an Associate Professor of Advanced Engineering in the department of Applied Electronics at the Tokyo University of Science (TUS), Japan. He received his Ph.D. from TUS Graduate School in 2004. His chief areas of interest are simulations of chemical compounds, biophysics, and protein folding simulations. A well-respected researcher, Dr. Ando has 42 publications to his credit.
About Professor Koji Tamura
from Tokyo University of Science
Funding information
This work was supported by the Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (JSPS) (Grant No. JP20K06592 to T.A. and Grant No. JP21K06293 to K.T.).

